How Would You Characterize The Transcription Obtained From The Control Template?
If DNA is a book, and so how is it read? Learn more about the DNA transcription process, where Deoxyribonucleic acid is converted to RNA, a more portable ready of instructions for the prison cell.
The genetic code is frequently referred to equally a "blueprint" considering it contains the instructions a cell requires in club to sustain itself. We now know that there is more to these instructions than simply the sequence of messages in the nucleotide code, however. For case, vast amounts of bear witness demonstrate that this code is the footing for the product of diverse molecules, including RNA and protein. Enquiry has as well shown that the instructions stored within DNA are "read" in ii steps: transcription and translation. In transcription, a portion of the double-stranded DNA template gives rise to a single-stranded RNA molecule. In some cases, the RNA molecule itself is a "finished product" that serves some of import function inside the cell. Often, even so, transcription of an RNA molecule is followed past a translation footstep, which ultimately results in the production of a protein molecule.
Visualizing Transcription
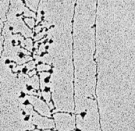
The procedure of transcription tin can be visualized by electron microscopy (Figure one); in fact, it was first observed using this method in 1970. In these early electron micrographs, the Deoxyribonucleic acid molecules appear as "trunks," with many RNA "branches" extending out from them. When DNAse and RNAse (enzymes that degrade DNA and RNA, respectively) were added to the molecules, the application of DNAse eliminated the trunk structures, while the use of RNAse wiped out the branches.
DNA is double-stranded, but only one strand serves as a template for transcription at any given time. This template strand is called the noncoding strand. The nontemplate strand is referred to as the coding strand because its sequence will be the same as that of the new RNA molecule. In most organisms, the strand of DNA that serves as the template for one cistron may be the nontemplate strand for other genes inside the same chromosome.
The Transcription Process
The procedure of transcription begins when an enzyme chosen RNA polymerase (RNA politician) attaches to the template DNA strand and begins to catalyze production of complementary RNA. Polymerases are big enzymes composed of approximately a dozen subunits, and when active on DNA, they are besides typically complexed with other factors. In many cases, these factors bespeak which gene is to be transcribed.
Three different types of RNA polymerase be in eukaryotic cells, whereas bacteria have just one. In eukaryotes, RNA politico I transcribes the genes that encode virtually of the ribosomal RNAs (rRNAs), and RNA politician 3 transcribes the genes for i small rRNA, plus the transfer RNAs that play a key role in the translation process, every bit well as other modest regulatory RNA molecules. Thus, it is RNA pol II that transcribes the messenger RNAs, which serve as the templates for production of protein molecules.
Transcription Initiation

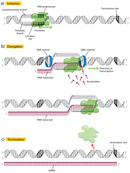
The outset step in transcription is initiation, when the RNA pol binds to the DNA upstream (five′) of the factor at a specialized sequence chosen a promoter (Figure 2a). In bacteria, promoters are usually composed of three sequence elements, whereas in eukaryotes, there are every bit many as 7 elements.
In prokaryotes, most genes have a sequence called the Pribnow box, with the consensus sequence TATAAT positioned nearly ten base of operations pairs abroad from the site that serves equally the location of transcription initiation. Not all Pribnow boxes have this exact nucleotide sequence; these nucleotides are but the most mutual ones institute at each site. Although substitutions do occur, each box nonetheless resembles this consensus adequately closely. Many genes also take the consensus sequence TTGCCA at a position 35 bases upstream of the start site, and some have what is called an upstream element, which is an A-T rich region 40 to lx nucleotides upstream that enhances the rate of transcription (Figure 3). In any instance, upon binding, the RNA politician "core enzyme" binds to another subunit chosen the sigma subunit to grade a holoezyme capable of unwinding the Deoxyribonucleic acid double helix in club to facilitate access to the gene. The sigma subunit conveys promoter specificity to RNA polymerase; that is, it is responsible for telling RNA polymerase where to bind. There are a number of unlike sigma subunits that bind to dissimilar promoters and therefore help in turning genes on and off as weather condition change.
Eukaryotic promoters are more circuitous than their prokaryotic counterparts, in part because eukaryotes have the aforementioned 3 classes of RNA polymerase that transcribe unlike sets of genes. Many eukaryotic genes also possess enhancer sequences, which can be found at considerable distances from the genes they bear upon. Enhancer sequences control gene activation past binding with activator proteins and altering the iii-D structure of the Dna to help "attract" RNA politico 2, thus regulating transcription. Because eukaryotic DNA is tightly packaged every bit chromatin, transcription also requires a number of specialized proteins that help brand the template strand accessible.
In eukaryotes, the "cadre" promoter for a factor transcribed by pol 2 is near often institute immediately upstream (5′) of the start site of the gene. Most politician II genes have a TATA box (consensus sequence TATTAA) 25 to 35 bases upstream of the initiation site, which affects the transcription rate and determines location of the showtime site. Eukaryotic RNA polymerases utilise a number of essential cofactors (collectively called full general transcription factors), and 1 of these, TFIID, recognizes the TATA box and ensures that the correct get-go site is used. Another cofactor, TFIIB, recognizes a unlike mutual consensus sequence, Grand/C G/C G/C Chiliad C C C, approximately 38 to 32 bases upstream (Figure 4).
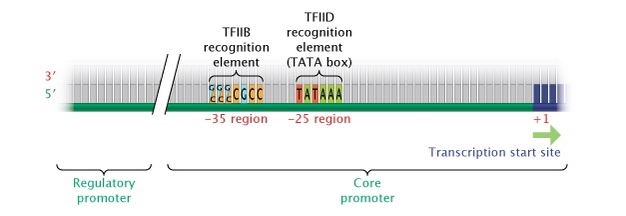
Figure four: Eukaryotic core promoter region.
In eukaryotes, genes transcribed into RNA transcripts by the enzyme RNA polymerase Two are controlled by a core promoter. A core promoter consists of a transcription start site, a TATA box (at the -25 region), and a TFIIB recognition element (at the -35 region).
© 2014 Nature Teaching Adapted from Pierce, Benjamin. Genetics: A Conceptual Arroyo, 2d ed. All rights reserved. ![]()
The terms "strong" and "weak" are often used to draw promoters and enhancers, co-ordinate to their effects on transcription rates and thereby on cistron expression. Amending of promoter strength can accept deleterious furnishings upon a cell, often resulting in disease. For example, some tumor-promoting viruses transform healthy cells by inserting strong promoters in the vicinity of growth-stimulating genes, while translocations in some cancer cells identify genes that should be "turned off" in the proximity of strong promoters or enhancers.
Enhancer sequences do what their name suggests: They act to heighten the rate at which genes are transcribed, and their furnishings tin exist quite powerful. Enhancers can be thousands of nucleotides away from the promoters with which they interact, simply they are brought into proximity by the looping of Deoxyribonucleic acid. This looping is the result of interactions between the proteins bound to the enhancer and those bound to the promoter. The proteins that facilitate this looping are called activators, while those that inhibit it are chosen repressors.
Transcription of eukaryotic genes by polymerases I and III is initiated in a similar mode, but the promoter sequences and transcriptional activator proteins vary.
Strand Elongation
Once transcription is initiated, the Deoxyribonucleic acid double helix unwinds and RNA polymerase reads the template strand, calculation nucleotides to the 3′ terminate of the growing chain (Figure 2b). At a temperature of 37 degrees Celsius, new nucleotides are added at an estimated rate of about 42-54 nucleotides per 2d in bacteria (Dennis & Bremer, 1974), while eukaryotes proceed at a much slower footstep of approximately 22-25 nucleotides per 2nd (Izban & Luse, 1992).
Transcription Termination
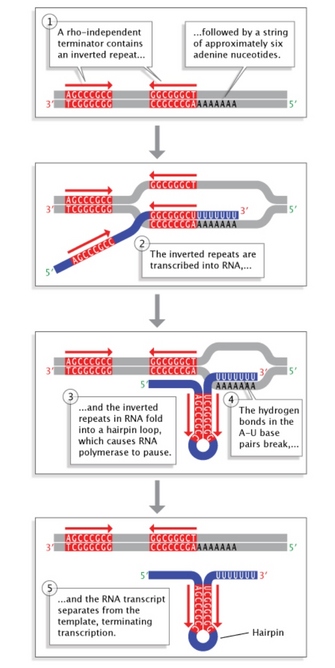
Figure 5: Rho-independent termination in leaner.
Inverted repeat sequences at the end of a gene permit folding of the newly transcribed RNA sequence into a hairpin loop. This terminates transcription and stimulates release of the mRNA strand from the transcription machinery.
© 2014 Nature Pedagogy Adjusted from Pierce, Benjamin. Genetics: A Conceptual Approach, 2d ed. All rights reserved. ![]()
Terminator sequences are constitute shut to the ends of noncoding sequences (Figure 2c). Bacteria possess two types of these sequences. In rho-independent terminators, inverted repeat sequences are transcribed; they tin can then fold back on themselves in hairpin loops, causing RNA pol to pause and resulting in release of the transcript (Figure 5). On the other hand, rho-dependent terminators make use of a cistron called rho, which actively unwinds the DNA-RNA hybrid formed during transcription, thereby releasing the newly synthesized RNA.
In eukaryotes, termination of transcription occurs by different processes, depending upon the verbal polymerase utilized. For political leader I genes, transcription is stopped using a termination factor, through a mechanism similar to rho-dependent termination in leaner. Transcription of pol III genes ends after transcribing a termination sequence that includes a polyuracil stretch, by a mechanism resembling rho-contained prokaryotic termination. Termination of pol II transcripts, however, is more circuitous.
Transcription of pol II genes can go on for hundreds or even thousands of nucleotides across the end of a noncoding sequence. The RNA strand is then cleaved past a complex that appears to associate with the polymerase. Cleavage seems to be coupled with termination of transcription and occurs at a consensus sequence. Mature political leader II mRNAs are polyadenylated at the 3′-end, resulting in a poly(A) tail; this process follows cleavage and is also coordinated with termination.
Both polyadenylation and termination make use of the same consensus sequence, and the interdependence of the processes was demonstrated in the late 1980s by piece of work from several groups. I group of scientists working with mouse globin genes showed that introducing mutations into the consensus sequence AATAAA, known to be necessary for poly(A) addition, inhibited both polyadenylation and transcription termination. They measured the extent of termination by hybridizing transcripts with the different poly(A) consensus sequence mutants with wild-blazon transcripts, and they were able to see a subtract in the indicate of hybridization, suggesting that proper termination was inhibited. They therefore ended that polyadenylation was necessary for termination (Logan et. al., 1987). Some other grouping obtained similar results using a monkey viral organization, SV40 (simian virus 40). They introduced mutations into a poly(A) site, which caused mRNAs to accrue to levels far in a higher place wild blazon (Connelly & Manley, 1988).
The exact relationship between cleavage and termination remains to be adamant. One model supposes that cleavage itself triggers termination; another proposes that polymerase activity is afflicted when passing through the consensus sequence at the cleavage site, perhaps through changes in associated transcriptional activation factors. Thus, research in the surface area of prokaryotic and eukaryotic transcription is yet focused on unraveling the molecular details of this circuitous process, data that will allow united states to ameliorate understand how genes are transcribed and silenced.
References and Recommended Reading
Connelly, Due south., & Manley, J. Fifty. A functional mRNA polyadenylation betoken is required for transcription termination past RNA polymerase II. Genes and Development 4, 440–452 (1988)
Dennis, P. P., & Bremer, H. Differential rate of ribosomal protein synthesis in Escherichia coli B/r. Periodical of Molecular Biology 84, 407–422 (1974)
Dragon. F., et al. A large nucleolar U3 ribonucleoprotein required for 18S ribosomal RNA biogenesis. Nature 417, 967–970 (2002) doi:10.1038/nature00769 (link to article)
Izban, M. K., & Luse, D. S. Factor-stimulated RNA polymerase Two transcribes at physiological elongation rates on naked DNA but very poorly on chromatin templates. Journal of Biological Chemical science 267, 13647–13655 (1992)
Kritikou, E. Transcription elongation and termination: Information technology ain't over until the polymerase falls off. Nature Milestones in Factor Expression 8 (2005)
Lee, J. Y., Park, J. Y., & Tian, B. Identification of mRNA polyadenylation sites in genomes using cDNA sequences, expressed sequence tags, and trace. Methods in Molecular Biology 419, 23–37 (2008)
Logan, J., et al. A poly(A) add-on site and a downstream termination region are required for efficient cessation of transcription by RNA polymerase Two in the mouse beta maj-globin factor. Proceedings of the National Academy of Sciences 23, 8306–8310 (1987)
Nabavi, S., & Nazar, R. N. Nonpolyadenylated RNA polymerase II termination is induced by transcript cleavage. Journal of Biological Chemistry 283, 13601–13610 (2008)
How Would You Characterize The Transcription Obtained From The Control Template?,
Source: https://www.nature.com/scitable/topicpage/dna-transcription-426/
Posted by: smithmisho1978.blogspot.com


0 Response to "How Would You Characterize The Transcription Obtained From The Control Template?"
Post a Comment